Started a github repository to put online R functions I've create for some common types of analysis and plots. Aim to have a core set of functions to make figures look prettier, even on preliminary analysis. A couple examples are included focusing on the first function: principal component analysis.
A GitHub repository can be found at:
R plotting functions.
For those who want to dive right in, the git repository (includes a readme):
R plotting functions
Update: The PCA script presented here could be greatly simplified using ggplot, a very useful graphics program in R. I'll leave implementation for a future post.
Been writing a lot of R functions and trying to make them generalizable and accepting of multiple input types. Since I was helped by others who posted code and other useful information about R online, thought I should contribute back.
There are a multitude of packages to help create useful and pretty plots with R. But sometimes it is also helpful to have functions that combine features of these packages into one nice example. That is what I hope to achieve with this repository. Each type of plot or analysis will have its own functions, examples and plots. This will allow users to verify that the functions work. Further, I hope those who just want to see a particular R feature implemented within a wider, working function can benefit from this.
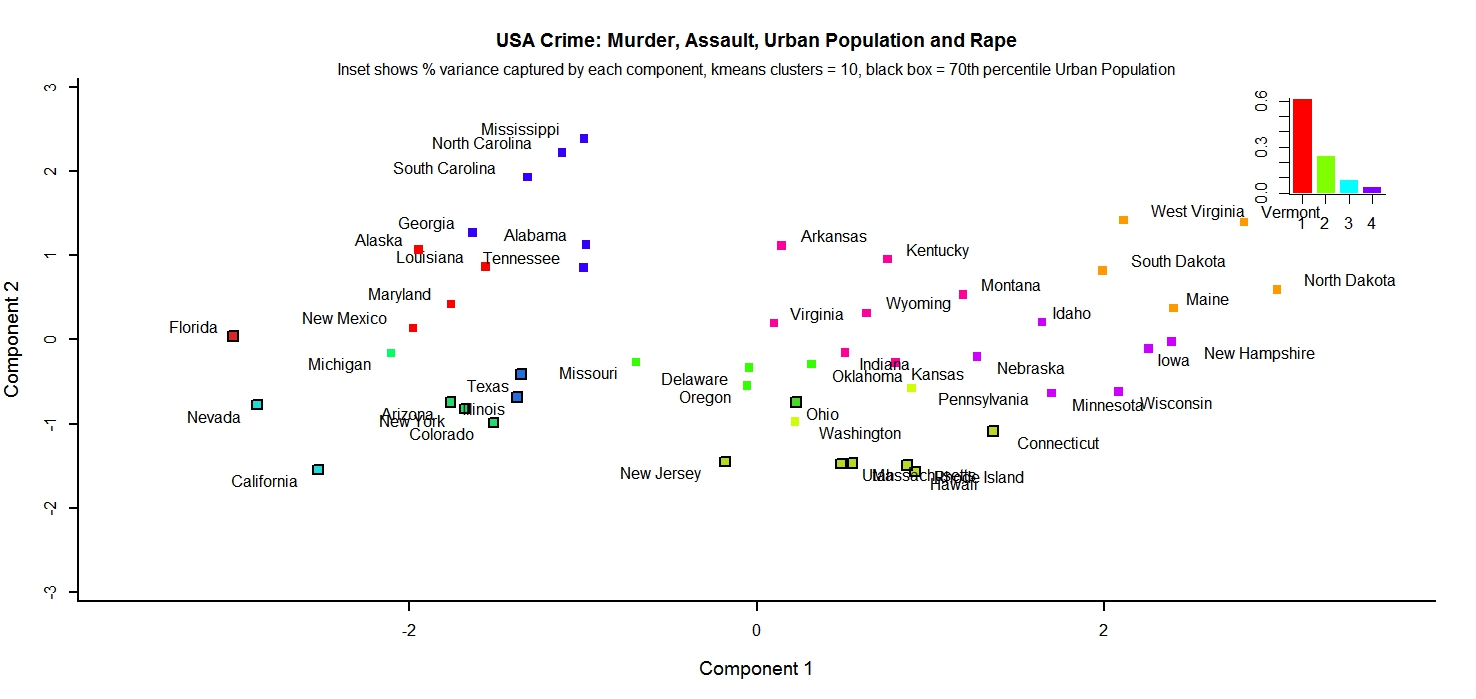
I have included two example images created with my first function, a script to do principal component analysis on arbitrary datasets. By getting the scores from the PCA object R creates, I can create a plot that is softer on the eyes that biplot or other standard functions. In addition, it allows me to input any arbitrary list and have the function highlight the subset of items in that list on the component graph. This allow easier visualization and understanding for human readers.
The first example looks at multiple crime statistics in the USA across states. Analyzing each individually might not tell us much about crime in the US at it relates to each state, but by doing PCA we see that there is some relation between these variables and that states with large urban populations group together, seen by looking at the clustering of the 70th percentile states.
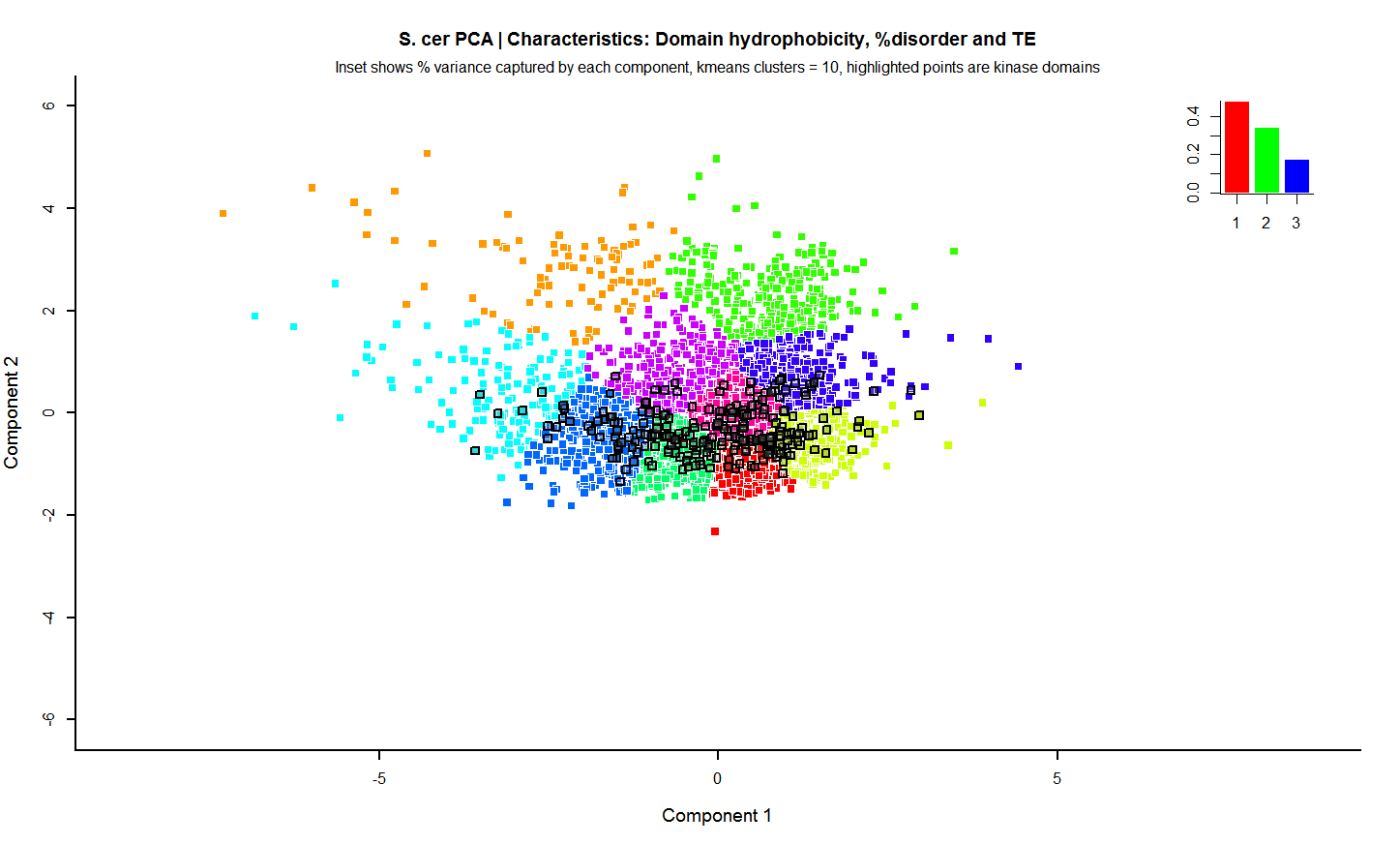
Next, I included some preliminary data, mostly uninformative to the uninitiated but visually nice, looking at biophysical protein properties across the entire yeast genome and then highlight the kinases to show this analysis can properly group related protein subsets. Obviously this is a rough first-analysis, but you get the picture.
Alright, this was supposed to be short, so I'll end it here. In the future I'll include code and explain the thought process behind it.
 stanford
stanford linkden
linkden github
github goodreads
goodreads medium
medium twitter
twitter